| Main | X-ray | Ligand analysis | Geometry details | Refinement progress | Real-space correlation coefficients | MolProbity analysis |
| All-Atom Contacts |
Clashscore, all atoms: | 10.39 | 88th percentile* (N=456, 2.20Å ± 0.25Å) | |
| Clashscore is the number of serious steric overlaps (> 0.4 Å) per 1000 atoms. | ||||
| Protein Geometry |
Poor rotamers | 11 | 3.56% | Goal: <1% |
| Ramachandran outliers | 3 | 0.88% | Goal: <0.05% | |
| Ramachandran favored | 321 | 93.86% | Goal: >98% | |
| MolProbity score^ | 2.36 | 62nd percentile* (N=10167, 2.20Å ± 0.25Å) | ||
| Cβ deviations >0.25Å | 0 | 0.00% | Goal: 0 | |
| Bad backbone bonds: | 0 / 1389 | 0.00% | Goal: 0% | |
| Bad backbone angles: | 0 / 1731 | 0.00% | Goal: <0.1% | |
For full MolProbity multichart with per-residue violation information, see here

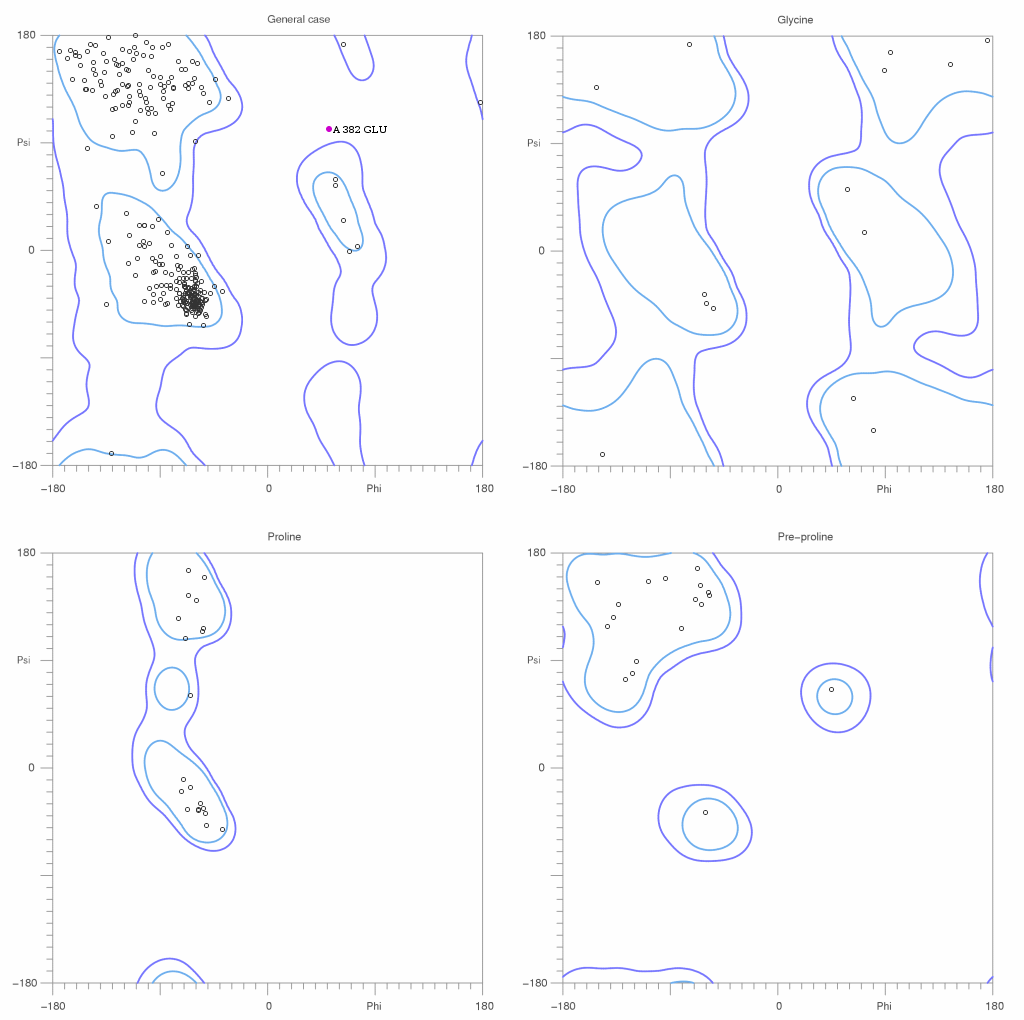
342 residues were evaluated in total for general, glycine, proline, and pre-pro.
95.61% of all residues were in favored (98%) regions. (327 residues)
99.71% of all residues were in allowed (>99.8%) regions. (341 residues)
There were 1 outliers:
A 382 GLU: (phi,psi) = (52.05, 100.76)
| Main | X-ray | Ligand analysis | Geometry details | Refinement progress | Real-space correlation coefficients | MolProbity analysis |